Get started
Geometry tools
Several espadon functions have been created to transform imaging volumes.
- Functions such as add.margin, nesting.cube, nesting.roi increase or decrease the number of voxels in the volume.
nesting.MR <- nesting.roi (MR, S, roi.sname = "brain", T.MAT = pat$T.MAT, vol.restrict = TRUE)
display.3D.stack (nesting.MR, ktext = FALSE, line.lwd = 2)
display.3D.contour (S, roi.sname = "brain", display.ref= MR$ref.pseudo, T.MAT = pat$T.MAT) 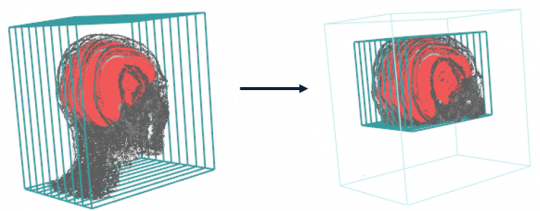
- The vol.regrid function resamples the volume to the grid of another imaging volume.
MR.on.CT <- vol.regrid (MR, CT, T.MAT = pat$T.MAT, interpolate = TRUE)
display.3D.stack (MR.on.CT, display.ref = CT$ref.pseudo , T.MAT = pat$T.MAT, ktext = FALSE, line.lwd = 2,
line.col = "#844A39") 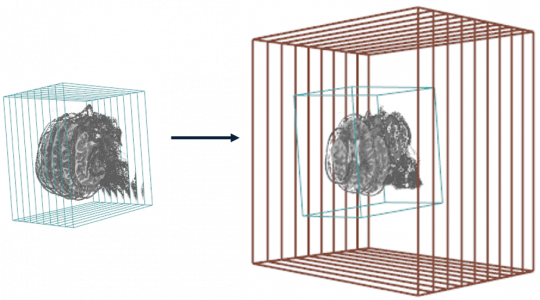
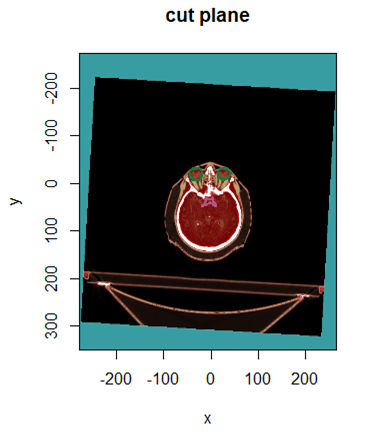
- More generally the get.plane function creates a cut of the volume in any direction. The get.direction function creates the direction vector from 3 points in space.
# Points determination : centers of the eyes and the chiasm found in rtstruct data
roi.idx <- select.names (S$roi.info$roi.pseudo, roi.name c("eye", "chiasm"))
pt <- S$roi.info[roi.idx, c("Gx", "Gy", "Gz")]
origin.pt <- as.numeric (apply (pt, 2, mean))
# Plane creation in the CT frame of reference
new.plane <- get.plane (CT, origin = origin.pt, plane.orientation = orientation.create (pt), interpolate = TRUE)
# Display of the new plane, in the cut plane frame of reference
tmat <- ref.cutplane.add (new.plane, origin.pt = origin.pt, ref.cutplane = "ref.plane", T.MAT = pat$T.MAT)
plan <- vol.in.new.ref (new.plane, "ref.plane", tmat)
display.plane (plan, struct= S, roi.idx = idx, T.MAT = tmat, view.coord = plan$patient.xyz0[3],
bottom.col = pal.RVV(200), bottom.breaks = seq(-1000, 1000, length.out = 201),
main = "cut plane", bg = "#379DA2", sat.transp = T, legend.plot = FALSE, interpolate = TRUE)